Search results for #MolecularNodes
We will be hosting a Reddit AMA session tomorrow, April 25th, at 12:00 PM USA ET to discuss our latest research published in @NatureMicrobiol . Please join us to ask your questions here: redd.it/1cbv9oj #Science #Discovery #MolecularNodes #STEAM nature.com/articles/s4156…
@nehaludyavar This particular animation was made using @Blender , an open source 3d software, Adobe Illustrator and #molecularNodes which is built and maintained by @bradyajohnston.
The argument for atheism is finished. Life is intelligible. This discovery has been published in @NatureMicrobiol #Explore #MolecularNodes #Discovery nature.com/articles/s4156…
The argument for atheism is finished. Life is intelligible. This discovery has been published in @NatureMicrobiol #Explore #MolecularNodes #Discovery nature.com/articles/s4156…
Another animation! This time looking at potassium ion transport across the cell membrane. Making good use of the amazing #MolecularNodes by @bradyajohnston to import molecular dynamics simulations directly into #Blender.
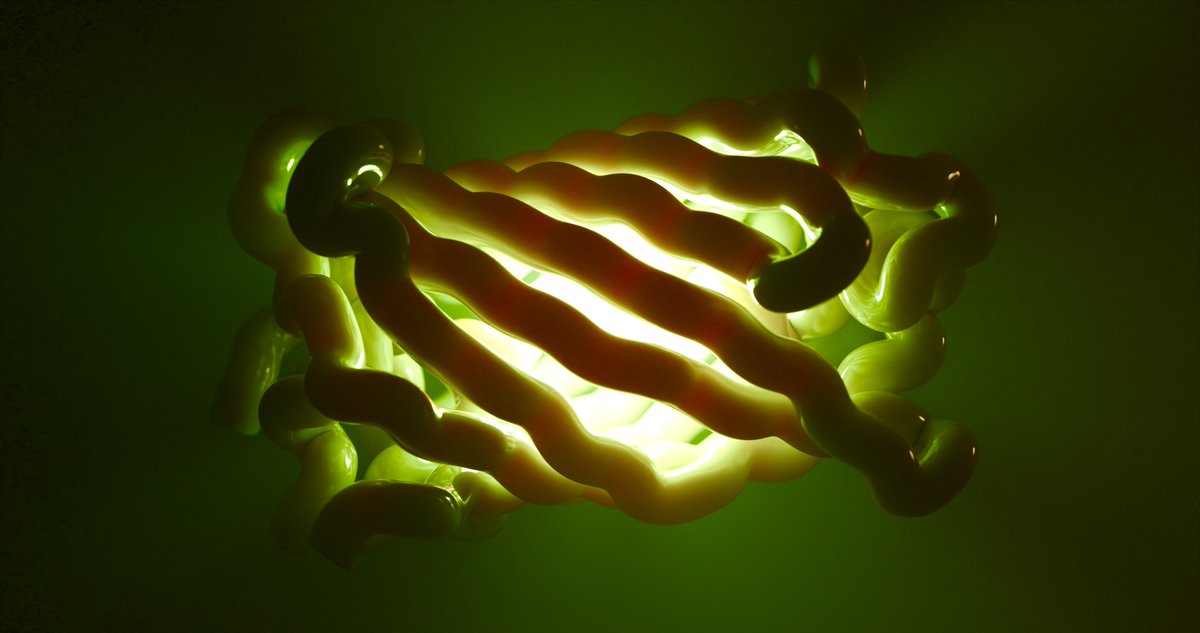
One of the most important proteins for experimental biology: GFP (work in progress) Making use of the intuitive selection and material assignment tools in @bradyajohnston 's #MolecularNodes to produce localised fluorescence. I think this still needs refinement...any thoughts?
The above structure render was created in @Blender using the #MolecularNodes add-on by @bradyajohnston and the CGRP peptide image was created using @UCSFChimeraX 👏
@prash_singh @NatureMicrobiol Amazing stuff @prash_singh , so great to see #MolecularNodes being used for animations like this 😍
دراسة تكشف آلية عمل المحرك البكتيري. (البروتينات في قاعدة سوط البكتيريا التي تفتل السوط الذي تستخدمه البكتيريا للحركة). @NatureMicrobiol #MolecularNodes #Discovery @prash_singh
Absolutely stunning CryoEM work on the flagellar motor from @prash_singh and company. Now I'm curious how the clutch mechanism works! Also, I love to see @bradyajohnston's #MolecularNodes out in the wild.
Absolutely stunning CryoEM work on the flagellar motor from @prash_singh and company. Now I'm curious how the clutch mechanism works! Also, I love to see @bradyajohnston's #MolecularNodes out in the wild.
@graveolens @NatureMicrobiol Hi Owen, the animation was made and rendered in @Blender, an open source 3D software using #MolecularNodes which is developed and maintained by @bradyajohnston.
@bradyajohnston @bradyajohnston inspired by your animation and help from your #MolecularNodes toolkit, here is an updated flagellar assembly with my newly discovered C-ring structure and symmetry mismatched MS-ring. #MolecularNodes made it feasible to render this complex animation on a laptop.
My grandfather built motors that powered big machines. In a lab not so different from his workshop, my colleagues and I uncovered the assembly of the bacterial motor. Today, this discovery has been published in @NatureMicrobiol #MolecularNodes #Discovery nature.com/articles/s4156…
For anyone interested I made a tutorial covering how to make the nanopore sequencing animation I posted last week😊 youtu.be/VV8b8A8_feM #b3d #MolecularNodes #sciart
Also, shoutout and much love to @bradyajohnston for fantastic Blender tutorials for protein structures. The title slide was made using his #MolecularNodes addon.
@MichaelSchwimm1 Everything looks great! I'm assuming the phospho stuff is done in Houdini. I'd be interested to get feedback on improvements to #MolecularNodes from those who have used houdini
Still more of a rough sketch than anything. But as I build better and more efficient lighting in Blender, @bradyajohnston's #molecularnodes just becomes even more powerful. Rendered using PDB 7JG7 and @DoubleGum_ 's incredible Clay Doh shader
Test animations of Nanopore Sequencing (ft. generic documentary music) from the Easter holiday. With significant help from @bradyajohnston's #MolecularNodes. Amazing what it can enable these days! #b3d #biology #sciart #sciencetwitter